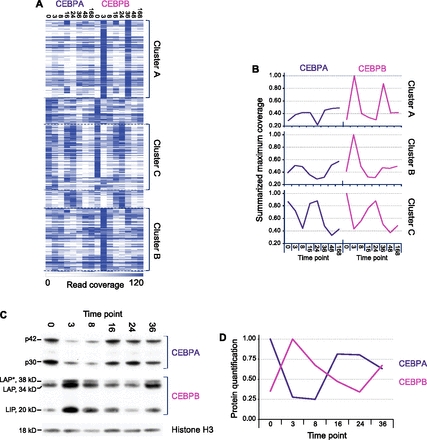
Three distinct temporal binding patterns reflect the CEBPA and CEBPB protein level ratios. (A) Heatmap of hierarchical clustering on CEBPA and CEBPB binding profiles, based on genomic coverage in each region (row) and time point (column). White signifies low binding level (coverage 0); dark blue, high (cut-off at coverage 120). The most prominent clusters are marked A, B, and C. (B) Summarized coverage profiles of the three clusters. For each profile, summed time point levels are shown as normalized to the highest point (set to 1). (C) Western blot showing CEBPA and CEBPB levels through the first 36 h of liver regeneration. Isoforms indicated (CEBPA: p42 and p30; CEBPB: LAP*, LAP, and LIP). Loading control is histone H3. Experiment is representative of three independent repeats. (D) Representative quantification of protein levels based on signal intensities (using ImageJ). All isoforms included for protein level determination, normalized to a maximum value of 1.











