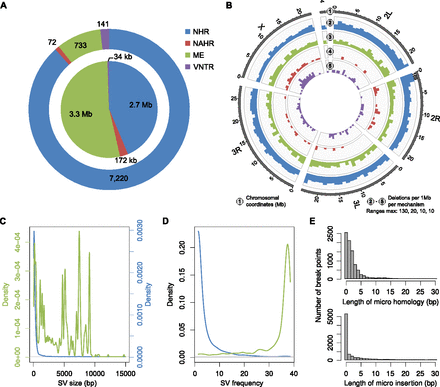
SV formation mechanisms acting in the Drosophila melanogaster genome. (A) Distribution of different SV formation mechanisms inferred amongst deletions for which the breakpoints were discovered at nucleotide resolution (8204 SVs). (Outer circle) Number of deletions per mechanism. (Inner circle) Cumulative size of these events. For 38 events the mechanism was ambiguous (Lam et al. 2010), and hence remained unclassified (data not shown in the plot). (B) Spatial distribution of deletions based on the different formation mechanisms. Colors as in A. In one case the bars exceeded the displayed range (thus, the absolute column height is indicated). (C) Size distribution of deletions related to NHR (blue) and ME (green). The peaks correspond to characteristic size spectra of mobile element classes active in the fruit fly. (D) Frequency distribution of deletions related to NHR (blue) and ME (green). A number of mobile elements were only present in the reference genome and were hence detected as deletions in the majority of samples. (E) Size distribution of micro homologies and micro insertions at deletion breakpoints.











