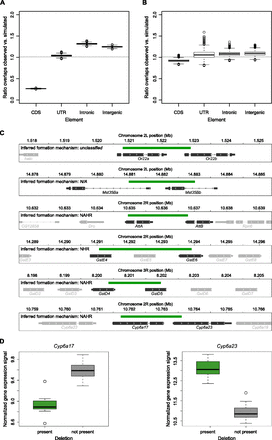
Functional impact of SVs. (A) Evidence for purifying selection against deletions overlapping gene coding sequences based on overlaps and expected overlaps with genomic annotation elements. The latter value was determined by randomly moving deletions within a 100-kb region and reanalyzing element overlaps 10,000 times. Intronic and intergenic deletions appeared slightly enriched relative to the randomized set, an effect that can be explained by the appreciable depletion of deletions overlapping coding sequence due to negative selection. (B) Analysis of tandem duplications overlapping annotated functional elements. (C) A collection of discovered putative fusion genes caused by partial gene deletion leading to hybrid genes. (Green) Deletions; (dark gray) fused genes; (light gray) genes in the vicinity of the locus in question. (CDS) Coding sequences; (UTR) upstream and downstream untranslated genic regions. (D) Gene expression analysis in male flies for Cyp6a17 and Cyp6a23 for samples with and without the gene fusing deletion that intersects with both genes. Variant categorization was achieved based on the genotype set (homozygous as well as heterozygous calls were considered as deletion).











