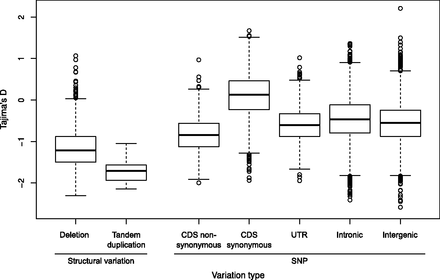
Genome-wide values of Tajima's D for SNPs and SVs. Boxplots depict genome-wide distributions of Tajima's D for SNPs in five genomic compartments (intergenic, intronic, nonsynonymous, synonymous, and UTRs), for deletions, and for tandem duplications, as estimated from sliding-window analyses. As classes, both deletions and tandem duplications show Tajima's D values significantly more negative than those of either synonymous or nonsynonymous SNPs (P < 1 × 10−8).











