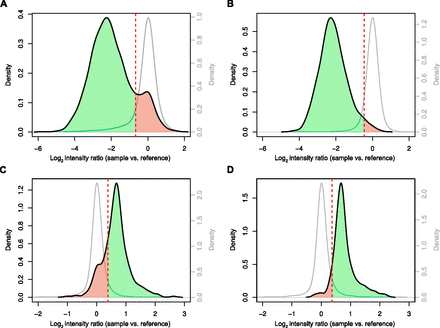
Whole-genome tiling array validation of the DGRP SV map. (A) Deletion discovery set. (Black line) Distribution of median log2 array intensity ratios for deletion loci, recorded between sample and the Berkley reference strain. (Gray line) Distribution for control samples lacking the SV at a given deletion locus of interest, which we used to determine the cutoff for FDR estimation (see Methods for details). (Dashed red line) Cutoff. SVs considered to be validated are highlighted in green, and potential false positive SVs are in red. (B) Deletion genotyping set. (C) Tandem duplication discovery set. (D) Tandem duplication genotyping set. Note that the different nature of homozygous deletions (causing a complete loss of genetic material) compared with homozygous duplications (causing mere duplication of material) likely render array-based FDR-estimations of duplications more error-prone (i.e., compare the better separation of hybridization-based curves for Fig. 3A,B compared with Fig. 3C,D for polymorphic vs. nonpolymorphic regions, and note the aforementioned PCR-based FDR estimates of 4%–6%).











