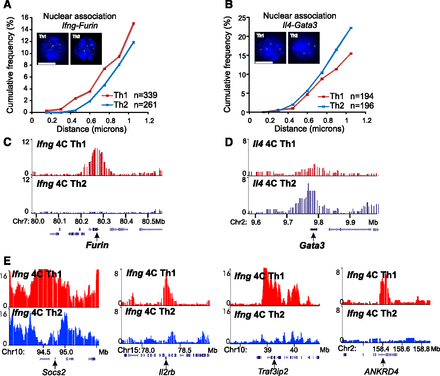
Cell type-specific interchromosomal association between functionally related genes in T lymphocytes. (A) 3D DNA FISH of Ifng and Furin in Th1 and Th2 lymphocytes. Representative projections of images and cumulative frequency of 3D distance are shown. Green foci indicate Ifng; red foci, Furin; and blue, DAPI staining. Scale bar, 5 microns. (n) Number of cells analyzed for Th1 (red curve) and Th2 (blue curve). (B) Similar analysis for the Il4 locus (green foci) and Gata3 (red foci). (C) Ifng 4C contact profiles at the Furin locus showing preferential contact in Th1 cells. (D) Il4 4C contact profiles at the Gata3 locus showing preferential contact in Th2 cells. (E) Th1-specific association of Ifng with Socs2, Il2rb, Traf3ip2, and ANKRD4 loci. Genomic coordinates in Mb (mm8). Th1 in red and Th2 in blue throughout the figures.











