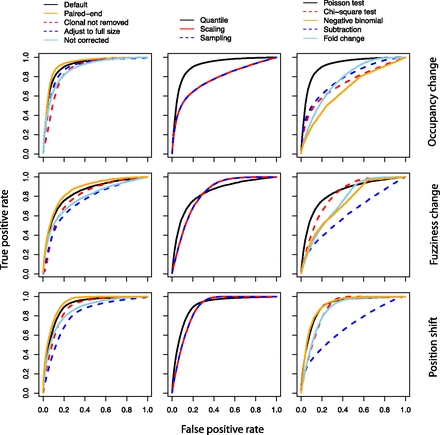
ROC curves indicate that the default DANPOS algorithm improves the identification of dynamic nucleosomes. Each column assesses the rate of true and false positive dynamic nucleosomes identified by different preliminary data processing methods (left), data normalization methods (middle), or statistical tests (right). Each row presents the results from simulation of nucleosome occupancy changes (top), fuzziness change (middle), and position shift (bottom). The known nucleosome changes were simulated from a nucleosome reference map (see Methods).











