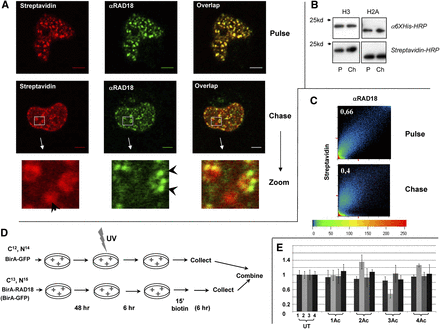
The RAD18-specific pattern changes after the proximity with RAD18 has been diminished. (A) Decrease in colocalization between RAD18 and biotinylated chromatin after a 6-h chase. (Top) Confocal microscopy showing strong colocalization between RAD18 and biotinylated chromatin in the nucleus of 293T cells immediately after pulse-labeling. (Middle) Confocal microscopy showing different localization of RAD18 and biotinylated chromatin 6 h after pulse-labeling. (Bottom) Zoomed area from the chase sample showing an example of biotinylated foci that do not colocalize with the RAD18 foci, and vice versa (indicated by arrows). (B) No increase in biotinylation signal after chase. Western analysis showing that the level of biotinylation of BAP-H3 and BAP-H2A is not increased 6 h after the cells have been washed free of biotin. (Top) α-6×His antibody. (Bottom) Strepavidin-HRP. (P) Pulse sample, (Ch) chase sample. (C) Scatter plot with the coefficient of correlation (top left corner) between the intensities of the red and green signals for every pixel (above a background threshold), showing stronger colocalization of the biotin and RAD18 signals in the pulse sample. (D) Scheme of the pulse-chase experiment. Cells grown in light SILAC medium are transfected by BirA-GFP and BAP-H2A (reference sample). Cells grown in heavy SILAC medium are transfected with BirA-RAD18 (or BirA-GFP) and BAP-H2A. The heavy labeled cells are either harvested immediately after biotin labeling or washed to remove biotin and left for 6 h before harvest. The heavy and light labeled cells are then mixed, and chromatin is prepared as in Figure 2C. (E) Acetylation status of H4 N-terminal tails. The analysis was performed and is presented as described in the legend of Figure 2D. The heavy isotope-labeled cells were: (lane 1) BirA-GFP + BAP-H2A pulse; (lane 2) BirA-GFP + BAP-H2A chase; (lane 3) BirA-RAD18 + BAP-H2A pulse; (lane 4) BirA-RAD18 + BAP-H2A chase. The light isotope-labeled cells were always transfected with BirA-GFP + BAP-H2A and pulse-labeled with biotin, as in sample 1.











