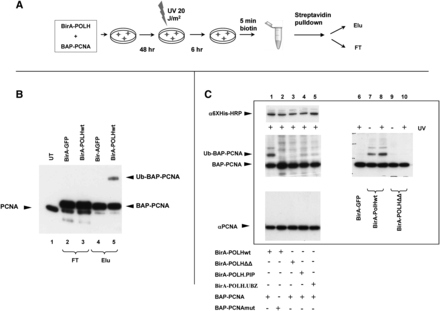
Analysis of post-translational modifications of a specific protein fraction using proximity utilizing biotinylation (PUB). (A) Experimental scheme. 293T cells are transfected with the plasmids expressing BAP-PCNA and BirA-POLH. Two days later, the cells are treated with UVC (20 J/m2), and 6 h afterward, the cells are pulse-labeled (5 min) with biotin. The cells are then harvested, and the biotinylated BAP-PCNA is separated from the nonbiotinylated BAP-PCNA using streptavidin-sepharose pulldown. (B) Ubiquitination status of POLH-proximal PCNA. Western blot with α-PCNA antibodies. (Lane 1) Untransfected cells; (lanes 2,4) BAP-PCNA cotransfected with BirA-GFP fusion; (lanes 3,5) BAP-PCNA cotransfected with Bir-POLH fusion; (lanes 2,3) flow-through fraction; (lanes 4,5) eluate. Note that the BAP-PCNA from the flow-through fraction was further purified via Ni-NTA chromatography in order to decrease the signal from endogenous PCNA. The endogenous PCNA, the BAP-PCNA fusion, and the ubiquitinated BAP-PCNA are indicated by arrows. (C) Ub-BAP-PCNA is preferentially biotinylated due to its specific interaction with POLH. Western blot analysis of nuclear extracts from transfected 293T cells. (Middle left and right panels) Strepatividin-HRP, detecting two forms of biotinylated BAP-PCNA in the presence of different Bir-POLH fusions, treated or untreated with UV; (top left) expression of different forms of BirA-POLH, detected by α-6×His Western blot; (bottom left) PCNA in the transfected 293 cells, showing that BAP-PCNA constitutes an undetectable fraction of the total PCNA in transfected cells. BirA-POLHwt, -.PIP, -.UBZ, -ΔΔ correspond, respectively, to wild-type POLH, mutant with deletion in the PCNA binding domain, point mutant D652A in the UBZ domain, and double mutant.











