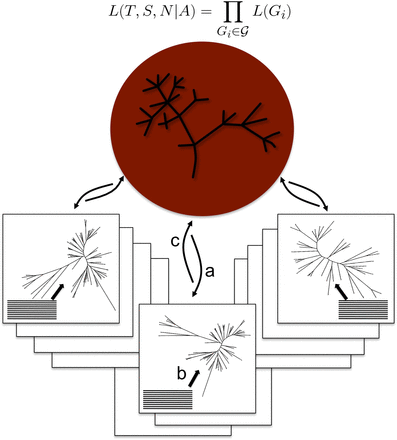
Genome-scale joint reconstruction of the species and gene trees. PHYLDOG is using a parallel server–client architecture. The server (in red) is in charge of the species tree search, and computes L(T, S, N|A). It communicates with clients (in boxes), each one in charge of one or more gene families, for which they search the gene tree maximizing L(Gi) = L(S, N|Ti) × L(Ti|Ai), using sequence alignments. (a) The server sends the current species tree as well as other parameters to the clients. (b) The clients compute L(Ti|Ai), i.e., the likelihood of a sequence alignment given the gene tree. ( c) The clients compute L(S, N|Ti), and send L(Gi) to the server.











