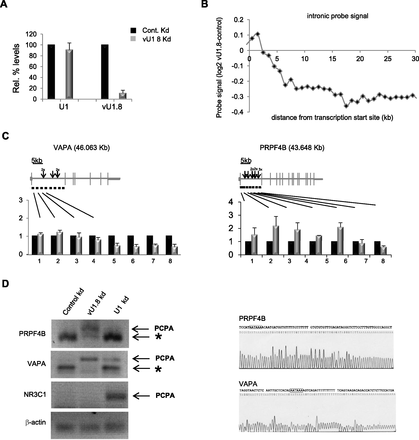
Altering vU1.8 snRNA levels cause global transcriptome changes in HeLa cells. (A) qRT-PCR analysis of vU1.8 and U1 snRNA levels in HeLa cells transfected with control or vU1.8 snRNA antisense oligonucleotides. (B) Average profile of intronic probe signal difference between vU1.8 snRNA knockdown and control cells across expressed nonoverlapping genes relative to the annotated transcription start sites. Each point represents an average of more than 100 probes in the corresponding region. (C) Validation of array results by qRT PCR, confirming the reduction in VAPA and PRPF4B pre-mRNA levels toward the 3′ end. Gene structures are illustrated above the graphs; horizontal and vertical lines indicate introns and exons, respectively. Location of primers is denoted as horizontal lines below the schematics. (Arrow) Cryptic poly(A) sites within the first intron of VAPA and PRPF4B genes. Sizes of genes are indicated in kilobases. (D) 3′ RACE, using nested PCR, on total RNA from HeLa cells transfected with control, vU1.8, or U1 snRNA antisense oligonucleotides was performed on the endogenous PRPF4B, VAPA, and NR3C transcripts to detect polyadenylated mRNAs. PCPA denotes a premature cleaved and polyadenylated mRNA product. (*) Mispriming of the oligo dT primer at an A-rich region. Primers targeting the endogenous beta-actin mRNA was used as a normalization control. Sequencing results of the PCPA products for the PRPF4B and VAPA gene are shown on the right. The putative poly(A) sites are indicated with black bordered boxes.











