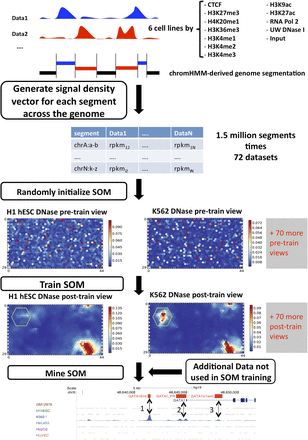
Training the self-organizing map and general overview of data analysis. The genome is first segmented based on the signal density of input data sets. Any segmentation approach can be applied; in this case, the ChromHMM-derived segmentation in the primary publications by The ENCODE Project Consortium was used. The signal density is calculated for each segment and each data set, resulting in an input matrix of M × N dimensions, where M is the number of segments and N the number of data sets. The SOM is then initialized randomly from the input matrix, and trained. Additional data sets, not used for training, can then be mapped to the SOM, and these mappings and the distribution of segments on the trained SOM can be mined for interesting biological relationships.











