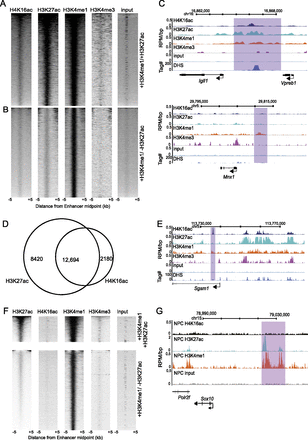
H4K16ac is found on active enhancers. (A,B) Heatmaps of H4K16ac, H3K27ac, H3K4me1, H3K4me3, and input ChIP-seq data (RPM) at H3K4me1+/H3K27ac+ (A), and H3K4me1+/H3K27ac− (B) marked enhancers. Data are ranked by sum of hits in the H4K16ac data set; 10-kb window around enhancer midpoint, in 500-bp windows. Intensity is determined by RPM. (C) Histone modification profiles and DNase I hypersensitive sites (DHS) across (top) a genetically defined enhancer active in ES cells (Igll1-Vpreb1 loci) and (bottom) a neuron-specific enhancer (at Mnx1 locus) not active in ESCs. Purple box indicates the likely enhancer location. Histone modifications are shown as RPM/bp in 200-bp sliding windows with 20-bp step, DHS sites are shown as tag density in a 150-bp window with a 20-bp step. (D) Quantification of SICER defined H3K27ac+/H3K4me1+/H3K4me3-, and H4K16ac+/H3K4me1+/H3K4me3- peak overlap. Venn diagram illustrates number of clustered active enhancers marked by H4K16ac or H3K27ac alone, or both together. (E) Profiles of H4K16ac, H3K27ac, H3K4me1, and H3K4me3, at an example of an H4K16ac+/H3K27ac-/H3K4me1+ putative active enhancer. Histone modifications and DHS sites shown as in C. (F) Heatmaps of H4K16ac, H3K27ac, H3K4me1, H3K4me3, and input ChIP-seq data (RPM) at H3K4me1+/H3K27ac+ (top), and H3K4me1+/H3K27ac− (bottom) marked enhancers in NPCs. Data are ranked by sum of hits in the H3K27ac data set; 10-kb window around enhancer midpoint, in 500-bp windows. Intensity is determined by RPM. (G) Profiles of NPC H4K16ac, H3K27ac, H3K4me1, and H3K4me3, as an example of an enhancer active in neuronal cell types (Antonellis et al. 2008). Histone modifications and DHS sites shown as in C.











