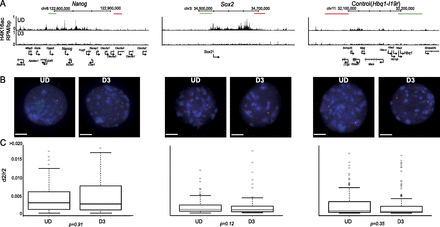
Loss of H4K16 acetylation does not correlate to chromatin compaction in vivo. (A) H4K16ac (RPM/bp in a 200-bp sliding window with a 20-bp step) across the Nanog, Sox2, and control (Hbq-Il9r) loci in undifferentiated ESC (UD, top row) and in differentiated cells (D3, bottom row). The position of fosmid probes (green and red boxes) used in FISH is indicated. Genomic maps are from the mm9 assembly of the mouse genome. (B) Example FISH images of nuclei from undifferentiated (UD; left) and differentiated (D3; left) ESCs, hybridized with probe pairs cross the Nanog, Sox2, and Hbq-Il9r loci. Nuclei were counterstained with DAPI (blue). Scale bar, 10 μm . (C) Boxplots indicating the distribution of squared interprobe distances (d2) normalized to nuclear radius2 (r2) for UD and D3 cells. Boxes show the median and interquartile range of the data; circles indicate outliers. n = 50 nuclei. Statistical significance of differences were examined by a Mann-Whitney U-test in R version 2.14.0.











