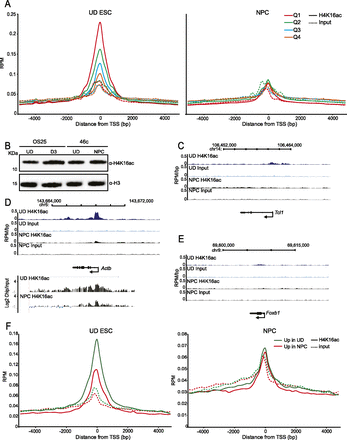
Gain and loss of H4K16 acetylation during ESC differentiation. (A) Normalized (average reads per million/RPM) H4K16ac (solid lines) or input (dotted lines) ChIP-seq tag counts around (±5 kb) the transcription start site (TSS) and transcription end site (TES) of genes separated into quartiles according to expression in ESC (left) or NPC (right) (from high to low [Q1–Q4]). (B) Western blot of H4K16ac in undifferentiated OS25 ESC, OS25 cells after 3 d of differentiation using retinoic acid (lanes 1 and 2), undifferentiated 46c ESC, and NPCs (lanes 3 and 4). Levels of H3 are shown for comparison. (C) UD H4K16ac, UD input, NPC H4K16ac, and NPC input profiles (RPM/bp), in 200-bp sliding windows with a 20-bp step, across the Tcl1 locus, which is silenced upon NPC differentiation. Exons are shown as boxes below the graph and the direction of transcription is indicated. (D) As in C, across the Actb locus, this maintains similarly high levels of expression in UD ESC and NPCs. At the bottom, log2 H4K16ac/input is shown at Actb established by hybridization to a custom microarray of ChIP DNA from 46c ESC and 46c NPC. Chromosome position and RefSeq gene annotations are used from July 2007 (mm9) mouse genome build (UCSC). (E) As in C, across the Foxb1 locus, this is silent in ESC, and highly expressed in NPCs. (F) RPM H4K16ac (solid lines) or input (dotted lines) ChIP-seq tag counts around (±5 kb) the TSS of genes differentially up-regulated in either ESC (ESC up genes, green lines) or NPC (NPC up genes, red lines). Data are shown for ESC (left) and NPC (right).











