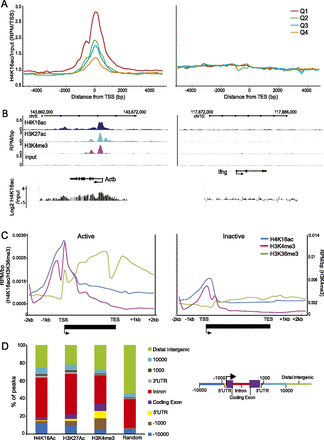
Profile of H4K16 acetylation in undifferentiated ESCs. (A) Input normalized (average reads per million/RPM) H4K16ac native ChIP-seq tag counts around (±5 kb) the transcription start site (TSS) and transcription end site (TES) of genes separated into quartiles according to expression in ESC from high to low (Q1–Q4). (B, top) H4K16ac/H3K27ac/H3K4me3 profiles (RPM/bp), in 200-bp sliding windows with a 20-bp step, across the Actb (active) and Ifng (silent) loci. Exons are shown as boxes below the graph, and the direction of transcription is indicated. (Bottom) Log2 H4K16ac/input at Actb and Ifng established by hybridization of H4K16ac ChIP DNA to a custom microarray. (C) Average RPM/bp tag counts across gene bodies for 500 active (left) and inactive (right) genes (in intervals of 10% of gene length) and extending 2 kb upstream of the TSS, and 2 kb downstream (in 200-bp intervals). Data are for H4K16ac (blue), H3K4me3 (pink), and H3K36me3 (green). (D) Genomic distribution of H4K16ac, H3K27ac, and H3K4me3 peaks in ESCs, and 0.5 × 106 peaks randomly distributed throughout the genome. (Left) Percentage of histone modification peaks found across each category of genomic sequence, relative to mm9 RefSeq genes. (Right) Schematic detailing the categorization (peaks classified as distal intergenic do not fall into any of the previous categories).











