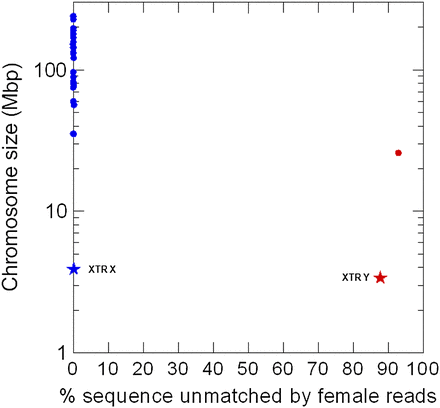
Test of the YGS method in the reference human genome. Each dot represents a human chromosome in the reference assembly (International Human Genome Sequencing Consortium 2004). (Blue dots) Autosomes and the X; (red dot) Y chromosome. The stars are the X-linked (blue star) and Y-linked (red star) copies of a 3.9-Mbp segmental duplication (“XTR”); the two copies are 98.8% identical (Ross et al. 2005). The region is continuous in the X chromosome (accession NT_011651.17; coordinates 11754754–15671817) and interrupted by a large insertion in the Y (Hughes and Rozen 2012); we used only the homologous region of the Y-linked copy (accession NT_011896.9; coordinates 268439–3453096 and 3751260–3967080). Note that the two copies of this segmental duplication are correctly identified as Y and not Y-linked, despite their very high sequence identity. (Abscissa) Proportion of scaffold sequence not matched by female short reads (in percentage unmatched single-copy k-mers); (ordinate) chromosome size.











