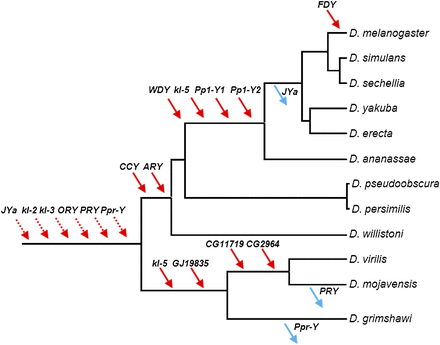
Gene gains and losses in the Drosophila Y. Gene location (Y-linkage vs. autosomal/X-linkage) was determined by PCR. Direction of the movements (gene gains, red arrows; gene losses, blue arrows) was inferred by synteny and parsimony (Supplemental Figs. S1–S4). For the six ancestral genes (dashed arrows), there is no close outgroup for inferring the direction (gain versus loss) (Koerich et al. 2008). Genes were labeled with the names of the D. melanogaster (JYalpha, CG11719, CG2964) or D. virilis (GJ19835) orthologs. Data for genes JYalpha (abridged as “JYa”), GJ19835, CG11719, and CG2964 came from the present study, and the remaining genes from Koerich et al. (2008).











