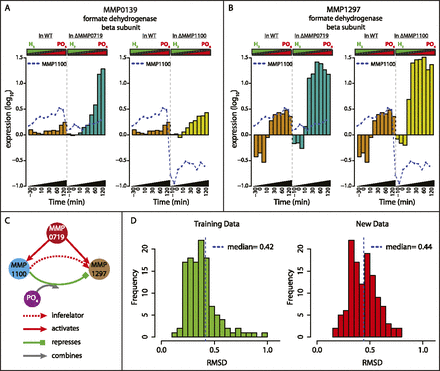
Roles of the transcriptional regulators MMP0719 and MMP1100. Transcriptional changes of formate dehydrogenase subunits in TF knockouts MMP1100 and MMP0719 during transition from P-limited to H2-limited conditions confirm EGRIN predictions. (A) barplot of fdhB2 (MMP0139) mRNA levels versus time in the wild-type (WT) (brown), MMP0719 knockout (dark green), and MMP1100 knockout (dark yellow) during transition from P-limiting to H2-limiting conditions. Expression of MMP1100 is also shown as blue dotted line in both genetic backgrounds. (B) Barplot of fdhB1 (MMP1297) mRNA changes, similar to A. (C) Based on the experimental confirmation of EGRIN predictions, we developed a model for the regulation of formate dehydrogenase. According to this model, MMP1100 modulates the activity of MMP1297 in a PO4-dependent manner, while MMP0719 activates both MMP1100 and MMP1297. (D) Predictive power of the EGRIN network model evaluated over the training (left) and new transcriptome data (right). Histogram of RMSD error between the predicted and measured response for 166 biclusters evaluated over 58 conditions in the training (left) and 56 conditions in the new data sets that were not used for the model construction (right). Similar median values (0.42 for training and 0.44 for new data set) indicate that our model performed equally well on both data sets.











