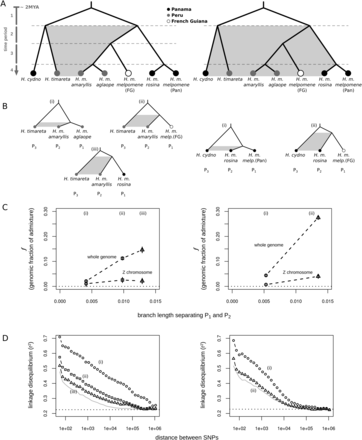
Measuring admixture at different phylogenetic scales. (A) We can distinguish between admixture in different time periods as follows. If gene flow was ancient only (i.e., period 1), then H. m. amaryllis and H. m. rosina should both be equally admixed with H. timareta and H. cydno. However, if gene flow is more recent (i.e., period 2, 3, or 4), then H. m. amaryllis should be more admixed with Peruvian H. timareta, and H. m. rosina should be more admixed with Panamanian H. cydno. The same logic applies when quantifying admixture for a specific branch: If H. timareta shares more derived alleles with H. m. amaryllis than with H. m. aglaope, this skew must reflect gene flow between H. timareta and H. m. amaryllis that is more recent than the coalescence between H. m. amaryllis and H. m. aglaope (i.e., during period 4). (B) Our sampling allowed us to quantify admixture at three time scales between H. timareta and H. m. amaryllis, and two time scales between H. cydno and H. m. rosina. (C) The estimated fraction of admixture (f), plotted for the whole genome and the Z chromosome specifically against the estimated length of the time period being analyzed, calculated as the average branch length separating populations P1 and P2 in the genomic ML phylogeny (Supplemental Fig. S1). Vertical lines depict standard errors. (D) LD (r2) between shared-derived alleles in the P2 population (left, H. m. amaryllis; right, H. m. rosina), plotted as a function of distance on a logarithmic scale. The SNPs used to estimate LD were those carrying a shared derived allele in P2 and P3, while P1 was fixed for the ancestral state (i.e., an ABBA pattern, where the B alleles are not necessarily fixed). The gray line represents the average genomic LD level, and the dashed line shows the average LD among unlinked sites.











