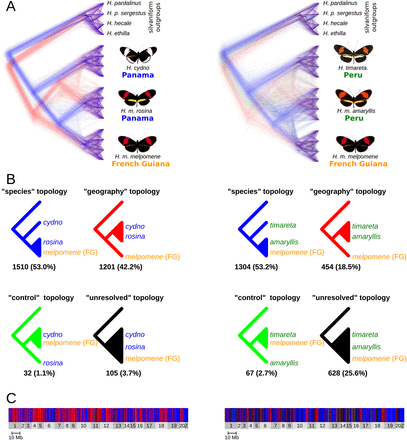
Four-taxon ML trees for 100-kb windows. (A) Trees were superimposed using DensiTree (Bouckaert 2010). There were 2848 trees for the H. cydno–H. melpomene data set (left) and 2453 for the H. timareta–H. melpomene data set (right). Tree lengths were equalized so that all trees could be superimposed, and then a random jitter was added to all branch lengths to show density. Trees supporting each of the four possible topologies are colored accordingly: blue for the species tree, red for the geography tree, green for the control tree, and black for unresolved trees. (B) The four topologies scored, along with the number and percentage of trees supporting each. See Supplemental Figure S3 for examples of trees assigned to each topology. (C) The distribution of the four topologies across the genome. Chromosomes are shaded light and dark gray. See Supplemental Figure S4 for an enlarged version.











