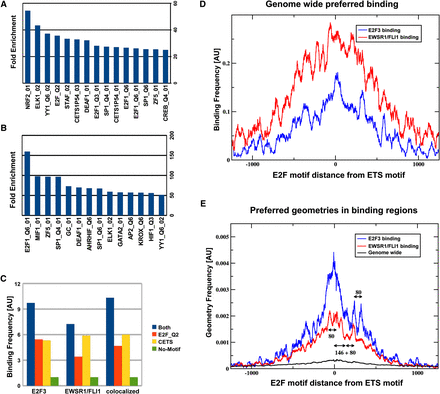
In silico analysis of binding regions. (A,B) Overrepresented transcription factor recognition sequences within binding regions of EWSR1/FLI1 and E2F3, respectively. The set of motifs shown in B has been selected in order to reduce redundancies between similar motifs. A complete list can be found in the Supplemental Material (Supplemental Table S3). (C) Binding frequency is correlated with the presence of ETS and E2F binding motifs in the promoter DNA sequence: Shown is the frequency of binding events of E2F3, EWSR1/FLI1, or both factors simultaneously (i.e., the average number of binding events per promoter). In this analysis promoters were subdivided into four groups, containing either E2F or ETS, neither, or both motifs. The group containing none of the two motifs was used for normalization, setting the frequency to one. Expanding the analysis to include the relative organization of ETS and E2F motifs in regions containing both motifs, it becomes apparent (D) that the effective binding affinity depends on that organization. The number of experimentally observed binding events (y-axis) overlapping any given ETS motif in the genome depends on the oriented distance (x-axis) from that ETS motif to the next E2F motif. (E) Specific spatial arrangements of ETS and E2F recognition sites are overrepresented in E2F3 and EWSR1/FLI binding regions. The plot displays the frequency of spatial configurations of ETS and E2F motifs within ChIP-seq binding regions of EWSR1/FLI1 and E2F3. Besides a global maximum at close distances, as expected in promoter regions with an overall increase of recognition sequences, discrete regions of increased frequency of pairs of E2F and ETS factors are visible. As a reference, scales for lengths 80 bp (length of one internucleosomal linker) and 146 bp (length of nucleosomal DNA) are also displayed in the graph. The frequency measurements depicted in D and E can best be interpreted as conditional probabilities, where D shows the probability to observe binding given that there is a specific geometric organization of the binding sites, while E shows the probability of finding a specific geometry, given that there is a binding event. Even though these two observations are related by Bayes' theorem, they are mutually independent, as discussed in detail in the Supplemental Material.











