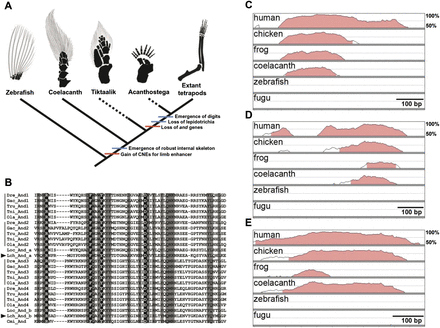
Genetic signature of fin-to-limb transition inferred from a genome comparison among vertebrate species. (A) Model of fin-to-limb transition based on morphological and molecular features. Red and blue bars indicate the molecular and morphological evolutionary events, respectively. The black and gray areas of the drawings depict the internal skeletons and lepidotrichia, respectively. The skeletons of the pectoral fins or limbs of zebrafish, Tiktaalik, Acanthostega, and mouse (extant tetrapods) were modified from Schneider et al. (2011). The skeleton of the pectoral fin of the coelacanth was drawn according to Millot and Anthony (1958). (B) Alignment of the N-terminal conserved domain of and genes showing two and genes in the coelacanth genome (arrowheads). Completely and mostly conserved (three or fewer amino acid substitution events during evolution) sites are shown with black and gray backgrounds, respectively. The species are indicated as follows: five teleost fishes, zebrafish (Dre: Danio rerio), stickleback (Gac: Gasterosteus aculeatus), fugu (Tru: Takifugu rubripes), pufferfish (Tni: Tetraodon nigroviridis), and medaka (Ola: Oryzias latipes); spotted gar (Loc: Lepisosteus oculatus); coelacanth (Lch: L. chalumnae); and elephant shark (Cmi: Callorhinchus milii). (C–E) VISTA plots of cis-regulatory elements in six vertebrate species using mouse as the reference for the following loci: (C) bmp7 intron 1 enhancer; (D) CNE11 in intron 10 of gli3; and (E) HMCO1 in the grem1-fmn1 locus. Lines indicate the degree of conservation from 50%–100%. The genomic regions estimated to be CNEs are shown by pink.











