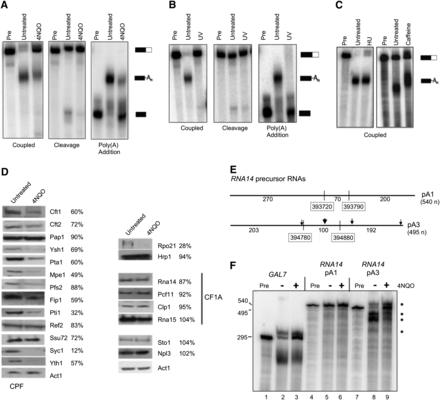
Effects of 4NQO on mRNA 3′-end processing. (A) In vitro processing efficiency. Yeast samples were treated with 5 μg/mL 4NQO for 2 h, followed by preparation of processing extracts. Coupled cleavage and poly(A) addition was assayed using radioactive precursor containing the GAL7 poly(A) site and flanking sequences (left panel). Cleavage was assayed by replacing ATP with dATP, which blocks tail extension (right panel, first three lanes). Poly(A) addition alone was assayed using RNA substrate ending at the GAL7 poly(A) (right panel, last three lanes). Unprocessed precursor is shown in the lanes marked “Pre,” and positions of substrate and product are shown on the right. (B) Effects of UV light on processing. Extracts were made from cells with or without exposure to UV light and examined for processing as described in A. (C) Effects of other agents on processing. Extracts were made from cells treated with hydroxyurea (HU, 10 mg/mL) or caffeine (20 mM) for 2 h, and coupled cleavage and poly(A) addition reactions compared with that of extract from untreated cells. Neither treatment inhibits the efficiency of processing, but we note that poly(A) tails are longer with exposure to caffeine. (D) Protein level variation. Western blots of Rpo21, processing complex subunits, Npl3, and Sto1, with actin (Act1) as the loading control. The percentage of protein remaining after 4NQO treatment was calculated after normalizing to Act1 and is indicated next to each protein. (E) Diagram of RNA14 substrates. This figure depicts the RNA14 precursors and indicates the position of poly(A) sites used in vitro in relationship to the region mapped for the upstream and downstream sites by DRS and shows the length of sequence in each portion of the transcript. The numbers marking the regions mapped by DRS correspond to the chromosomal positions shown in the CPD plot of Figure 2A. The product indicated for pA1 refers to the reactions in F. (F) Coupled cleavage and poly(A) addition reactions using the transcripts containing the first (pA1) or third (pA3) RNA14 poly(A) sites. Reactions were performed with GAL7 substrate (lanes 1–3), RNA14 pA1 (lanes 4–6), and RNA14 pA3 (lanes 7–9) using extract from cells with (+) or without (−) 4NQO treatment. Unprocessed precursor is shown in the lanes marked “Pre.” The products from the RNA14 pA3 substrate are indicated by an asterisk.











