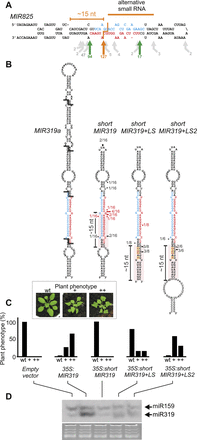
Switching structural determinants in plant miRNA precursors. (A) Scheme of MIR825. Cuts releasing the miRNA/miRNA* are indicated by the green arrows. Gray arrows show other less abundant cleavage sites. A frequent cut located at 15 nt of a lower stem is indicated with an orange arrow, as well as an alternative small RNA detected at low frequency. (B) Scheme of MIR319a precursor and mutants with a deleted terminal region and the addition of a structured lower stem. Structured regions in the lower stems are highlighted with pink boxes. The cleavage sites analyzed by 5′ RACE PCR are indicated by red and black lines. The length of the arrows and the number of sequenced clones indicate the relative cloning frequency of the intermediates. (C) Phenotypes caused by overexpression of the wild-type and mutant MIR319 precursors. Note that wild-type leaves are flat but become crinkled when miR319 is overexpressed. (Inset) Transgenic plants were classified according to the shape of their leaves. At least 50 independent transgenic plants were scored in each case. (D) Small RNA blots of transgenic Arabidopsis inflorescences expressing the different precursors. Each sample is a pool of 25 independent transgenic plants. The miRNAs are indicated in red and the miRNAs* in blue.











