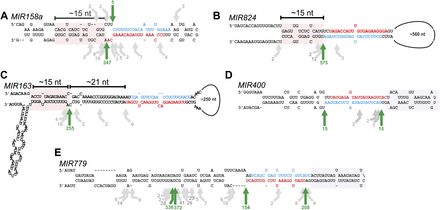
Processing of young miRNAs. Scheme showing the precursors of MIR158a (A), MIR824 (B), MIR163 (C), MIR400 (D), and MIR779 (E). The arrows indicate the positions and number of reads of the precursor cuts identified. Green arrows show the most abundant cleavage site detected, which also matches to the proximal and distal sides of the miRNA/miRNA*. Gray arrows show other cleavage sites that are less abundant. Structured regions in the lower stem (MIR158a, MIR824, and MIR163) and the upper stem (MIR400 and MIR779) are highlighted with pink and gray boxes, respectively. The miRNAs are indicated in red and the miRNAs* in blue.











