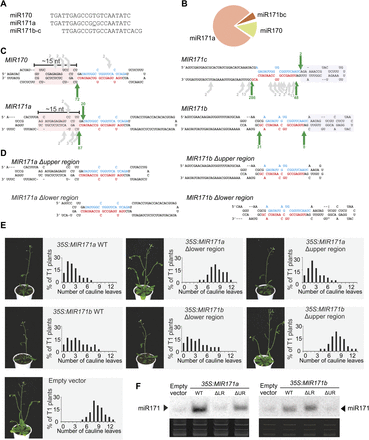
Mixed processing of MIR170/171 family members. (A) Sequences of miR170/171 miRNAs. (B) Relative abundance of the small RNAs by deep-sequencing small RNAs from wild-type plants. (C) Scheme showing the secondary structure and cleavage sites detected in the family members. Structured regions in the lower stem (MIR170, MIR171a) and upper stem (MIR171b/c) are highlighted with pink and gray boxes, respectively. The most abundant cleavage sites detected, which also release the miRNA/miRNA*, are indicated by green arrows. Gray arrows show other cleavage sites that are less abundant. The miRNAs are indicated in red and the miRNAs* in blue. (D) Scheme of MIR171a/b mutant precursors. (E) Phenotypes of transgenic Arabidopsis plants transformed with wild-type and mutant MIR171a/b precursors under the control of the 35S promoter. A representative transgenic plant overexpressing each construct is shown on the left. The panels show the cauline leaves' number distribution observed in at least 50 independent transgenic plants overexpressing each construct. (F) Small RNA blots showing the accumulation of miR171 in transgenic Arabidopsis seedlings expressing the different mutant precursors. (UR) Upper region; (LR) lower region. Each sample is a pool of at least 20 independent transgenic plants.











