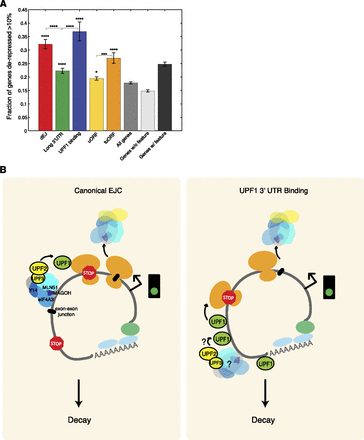
NMD features and models of UPF1-dependent mRNA repression. (A) Predictive capacity of mRNA features for NMD-regulation. Fractions of expressed genes harboring a dEJ (red), long 3′ UTR (green), UPF1 3′ UTR binding (blue), uORF (light orange), and/or tuORF (orange) that were derepressed consistently are shown. Shown for comparison are the fractions of genes derepressed consistently regardless of feature content (medium gray), without any NMD-inducing feature (light gray), and with at least one feature (dark gray). P-values above each feature indicate significance relative to all expressed genes; brackets indicate significant comparisons between features (hypergeometric test). See also Supplemental Figure S5A. Asterisks as in Figure 1. (B) (Left) Canonical dEJ-mediated regulation. EJC components (blue and gray) are deposited as a consequence of splicing in the nucleus ∼24 nt upstream of an exon-exon junction (black bar). Members of the EJC, including UPF2 and UPF3, help to stabilize the transient UPF1-ribosome interaction as well as to stimulate UPF1's phosphorylation and helicase activity, ultimately leading to decay of the message. (Right) 3′ UTR UPF1 binding-mediated regulation. UPF1 binds to mRNA 3′ UTRs independent of the presence of an exon-exon junction. At some frequency, UPF1 is activated by interaction with cytoplasmic EJC components. These factors may either be recently released from mRNAs due to translation or perhaps stably associated with message 3′ UTRs independent of an exon-exon junction.











