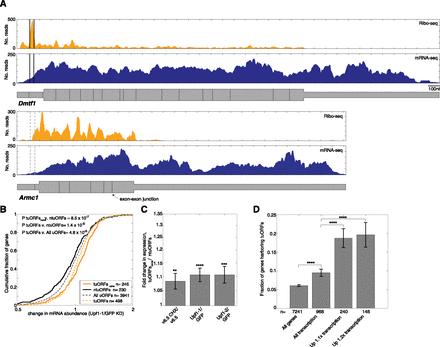
Translation of uORFs is associated with UPF1-mediated repression. (A) mRNA-seq (blue) and ribosome footprint (orange) reads from UPF1 depleted (shRNA Upf1-1) cells mapping to Dmtf1 and Armc1 mRNAs. Dmtf1 (top) contains a tuORF (outlined with dark gray lines), while Armc1 contains a single ntuORF (dashed lines). (B) CDFs of changes in mRNA abundance following UPF1 depletion (shRNA Upf1-1) for all genes with a tuORF (dashed yellow line), consistent genes with a tuORF (orange line), genes with a uORF (dashed gray line), and genes with an ntuORF (black line). P-values determined by Wilcoxon rank sum test. (C) Ratios of median fold expression change following NMD inhibition of consistent genes with a tuORF to genes with an ntuORF. Error bars represent standard error of the two populations compared. P-values determined as in B. (D) Fraction of genes harboring a tuORF for all expressed genes, all expressed transcriptional regulators (GO:0045449), and transcriptional regulators derepressed by more than 1.1- or 1.2-fold in at least two out of three NMD inhibition experiments. P-value of enrichment determined by hypergeometric test. Numbers of genes in each category are indicated. Error bars indicate binomial standard deviations. Fold change values and ratios shown in B and C are plotted on log2 scale. Asterisks as in Figure 1.











