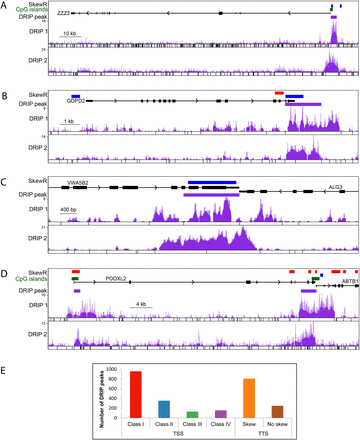
DRIP-seq illustrates R-loop formation at the 5′ and 3′ ends of human genes. (A–D) DRIP-seq profiles. The SkewR track shows regions of GC skew with red indicating G-rich blocks; blue, C-rich blocks. DRIP 1 and DRIP 2 correspond to DRIP-seq experiments for which the genome was fragmented with two distinct cocktails of restriction enzymes (cut sites are indicated below each DRIP data set). The DRIP peak track indicates the consensus DRIP signal. (A,B) An R-loop at the TSS and TTS of a gene, respectively. (C) R-loop forms at the TTSs of two convergent genes. (D) The PODXL2 gene shows both TSS and TTS R-loops. Note that the TTS is followed closely by the TSS of the neighboring ABTB1 gene. (E) Distribution of DRIP-seq peaks over TSS and TTS classes.











