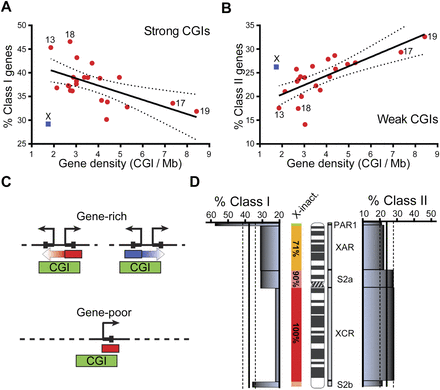
Gene density strongly affects the distribution of Class I and Class II genes and the X chromosome represents an exception to the autosomal trends. (A,B) The distribution of Class I and Class II genes on individual chromosomes is represented as a percentage of total RefSeq genes on that chromosome (y-axis) plotted against a measure of gene density (x-axis; CGI/Mb, a set of 10,279 high confidence promoter CGIs, was used) (Bock et al. 2007). The X chromosome is shown in blue; autosomes are in red; a few relevant chromosomes are indicated. The data was fit to a linear regression shown here with the corresponding 95% confidence interval. (C) Schematic representation of the manner by which a gene-rich region may enable a shared epigenetic state (arrows) between neighboring genes, while a gene-poor region may not. CGI promoters are shown by green boxes; peaks of G-skew or C-skew are shown by red and blue boxes, respectively. (D) The distribution of Class I (left) and Class II (right) genes is represented as a percentage of total RefSeq genes calculated over each X-chromosome evolutionary strata (PAR1, 0–2.8 Mb; XAR, 2.8–46.8 Mb; S2a, 46.8–60 Mb; XCR, 60–148.6 Mb; and S2b, 148.6–154.8 Mb). The expected percentage of Class I and Class II genes based on their autosomal distributions is shown by a straight line together with standard deviation (dotted lines). The X-inactivation efficiency across each strata is color-coded and was determined from Carrel and Willard (2005).











