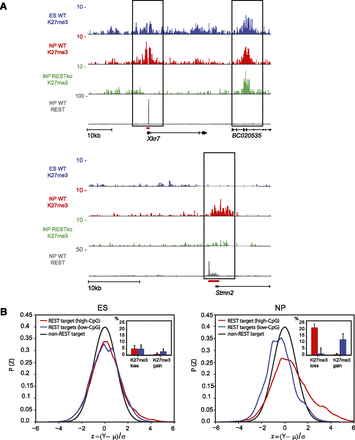
REST is required for H3K27me3 dynamics in NP cells. (A) ChIP-Seq signal for H3K27me3 and REST in representative genomic regions. Shown are H3K27me3 signal in ES cells, NPs of wild-type (WT) and RESTko cells, as well as REST signal in NPs. The top panel exemplifies selective loss of H3K27me3 at the REST binding site of the Xkr7 locus, whereas neighboring regions (BC020535) remain unaffected. The lower panel shows similar loss of H3K27me3 at the Stmn2 locus. Both the Xkr7 and Stmn2 loci are examples of promoter-proximal high-CpG regions. Shown are normalized read densities. The red bars at the REST peaks indicate the regions cloned for transgenic experiments. (B) Global comparison of H3K27me3 levels between wild-type and RESTko cells. Shown are the normalized distributions (see Methods) of the ratio between H3K27me3 in wild type versus RESTko for nontarget regions (black lines) and for either low-CpG (blue lines) or high-CpG (red lines) regions that are REST targets at the ES (left panel) and NP (right panel) stage. (Insets) Estimated fractions of REST targets that significantly lose or gain H3K27me3 in the RESTko at high-CpG (red) and low-CpG regions (blue). There are few significantly changing targets at the ES stage. At the NP stage a significant fraction of high-CpG targets lose H3K27me3 and a smaller but still significant fraction of low-CpG targets gain H3K27me3 in the RESTko cells.











