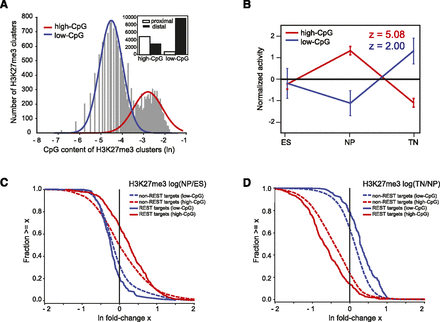
REST is associated with H3K27me3 dynamics at high- and low-CpG regions genome-wide. (A) The distribution of CpG dinucleotide frequencies of H3K27me3 regions genome-wide is bimodal and can be fit by a mixture of two log-normal distributions (red and blue lines) corresponding to high- and low-CpG regions, respectively. (Inset) Numbers of K27me3 regions that are promoter-proximal and distal for high-CpG and low-CpG regions. (B) REST activity profiles on high- (red) and low-CpG regions (blue) as inferred by running Epi-MARA on all H3K27me3 regions genome-wide show a transient gain and loss, respectively, at the NP stage. Note that, whereas REST activity on the high-CpG regions is highly significant, on the low-CpG regions REST activity has a much weaker significance. (C) Reverse cumulative distributions of changes in H3K27me3 levels at the transition from ES to NP stage. We divided regions that were enriched for H3K27me3 into high-CpG/low-CpG (red/blue) and REST-target/nontarget (solid/broken lines) regions. At high-CpG regions REST targets tend to gain H3K27me3 going from the ES to the NP stage whereas nontarget regions are equally likely to gain or lose H3K27me3. In contrast, most low-CpG regions lose H3K27me3 going to the NP stage, and REST targets tend to lose even more H3K27me3. (D) As in panel C but now for the transition from the NP to the TN stage. High-CpG regions generally tend to lose H3K27me3 and REST targets tend to lose even more, whereas low-CpG regions tend to gain H3K27me3 and REST targets tend to gain even more.











