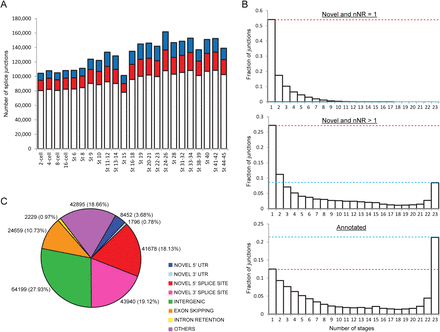
Novel splice junctions of different types can be detected at all developmental stages. (A) Total number of splicing events identified at every stage. While the majority of the junctions detected at each stage are annotated in either RefSeq or Ensembl (white portions), a sizeable number of junctions are novel (colored portions). (Red) There are two or more nonredundant reads supporting the novel junction (nNR > 1). (Blue) There is only one nonredundant read supporting the novel junction (nNR = 1). (B) Plots showing the number of stages that each splice junction is detected in. Compared to the novel junctions, a greater fraction of annotated junctions is constitutively expressed. On average, each annotated junction was detected in 11.0 stages, while each strongly supported novel junction (nNR > 1) was detected in 7.7 stages and each weakly supported novel junction (nNR = 1) was detected in 2.2 stages. (Red dotted line) Height of the bar for single-occurrence junctions; (blue dotted line) height of the bar for constitutive junctions. (C) Pie chart showing the different types of novel splice junctions identified with the number of junctions (percentage in brackets) given next to each segment. Only 27.93% of the novel junctions occur in intergenic regions (in green), while the majority of them (72.07%) occur within annotated genes.











