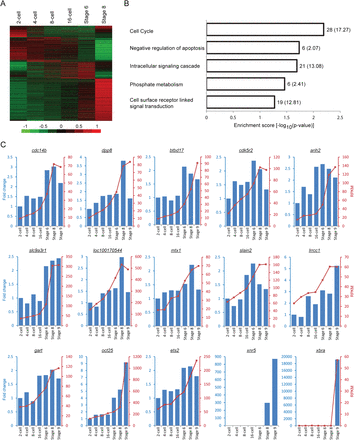
Many genes are transcribed before the midblastula transition despite a general state of transcriptional repression. (A) Heatmap showing distinct expression profiles of all RefSeq genes that were detected before the midblastula transition in our RNA-seq experiments. To focus on the stages before embryonic genome activation, only the RPKM values from the two-cell stage to Stage 8 were used for mean-centering, normalization, and clustering. Each row in the heatmap represents a different gene. (B) GO analysis of the 150 early transcribed genes that passed the following three criteria: (1) the transcript level at Stage 6 is at least twofold higher than that at the two-cell stage; (2) the average RPKM at each developmental stage is at least 1; and (3) the expression profile is monotonically increasing. (C) Validations of the RNA-seq results. The RT-qPCR expression profiles (blue bars) match the RNA-seq data (red lines) closely for 13 out of the 20 genes we tested. The nodal-related gene, xnr5, serves as a positive control (its expression is activated before EGA), while xbra serves as a negative control (its expression is activated after EGA). There is no RNA-seq data for xnr5 because it is duplicated in the Xenopus genome, and the corresponding reads cannot be uniquely mapped. We also note that for the majority of the validated genes, the fold changes obtained from our RT-qPCR experiments are relatively small (two- to threefold) compared to the fold changes observed for xnr5 and xbra (greater than 100-fold).











