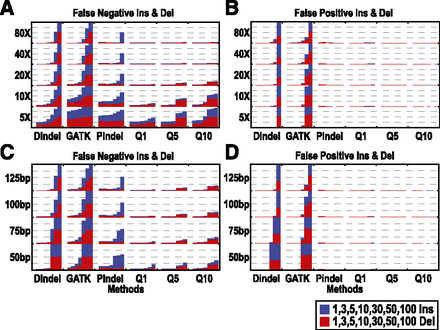
The effect of indel size, coverage, and read length on the false-negative and false-positive rates of SOAPindel, Dindel, Pindel, and GATK on simulated data based on hg19, chromosome 22. (A,B) FN and FP percentage for read length of 100 bp and coverage of 5/10/20/40/80×, with fixed indel sizes of 1/3/5/10/30/50/100. Every column is split into two parts: % of insertion and % of deletion. (C,D) FN and FP percentage for read lengths of 50/75/100/125 bp and coverage of 20×, with the same fixed indel sizes and also split into insertion and deletion.











