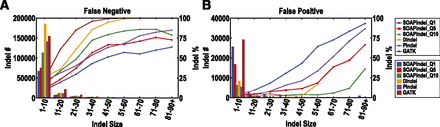
False-negative (FN) (A) and false-positive (FP) (B) rates of indels called by SOAPindel, Dindel, Pindel, and GATK on data simulated based on hg19. The read length is 100 bp, and the coverage is 20×, and SNP and indel variation are from the empirical differences of the Venter and hg19 genomes. The numbers of FP and FN indels refer to histograms, whereas the lines correspond to the percentage of indels being either FN or FP (secondary y-axis).











