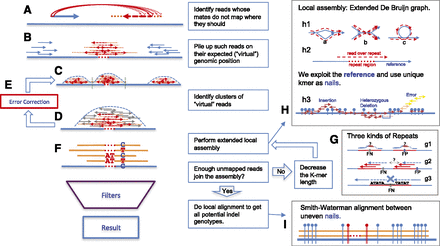
Figure 1.
Outline of the SOAPindel algorithm. (A) Identify nonmapping mates. (B) Pile up unmapped reads. (C) Identify clusters of unmapped reads. (D) Cut out clusters and add mapped reads. (E) Error correct clusters. (F) Find candidate indels by de novo assembly. (G) Treatment of repeats. (H) Extended local assembly with nails. (I) Local alignment between nails. For details, see Methods.











