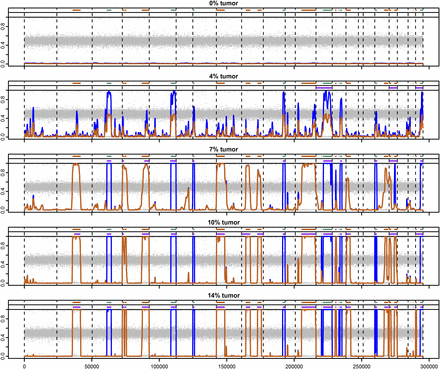
Local posterior probabilities of allelic imbalance at various dilutions. These whole-genome results are from a three-state HMM, accommodating two levels of imbalance. Horizontal lines at the top of each plot show the locations of simulated deletions (orange) and cn-LOH (green). Below these, purple bars show the regions identified by BAFsegmentation to contain AI. Vertical axes range from 0 to 1 for both the BAFs (gray points) and posterior probabilities (orange for deletions, blue for deletions and cn-LOH combined).











