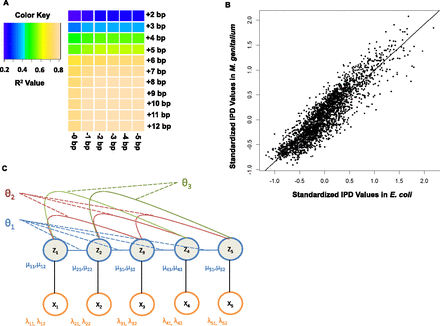
DNA polymerase kinetics in SMRT sequencing is a function of the local sequence context of the incorporation site, motivating
a conditional random field approach to KVE detection. (A) Heatmap of the coefficient of determination (R2) for the IPD variance for the incorporation site of a SMRT sequencing reaction explained by local sequence context. This
heatmap suggests that seven bases upstream of and two bases downstream from the incorporation site are the most informative,
and that bases beyond this context do not provide much additional information about the enzyme kinetics. (B) Scatter plot comparing IPDs in identical sequence contexts between whole-genome amplified E. coli and M. genitalium samples. Each point represents the log of the IPD for a given 10-bp context (seven bases upstream of and two bases downstream
from the incorporation site) in E. coli (y-axis) and M. genitalium (x-axis): 2500 points sampled from the 1,048,576 possible 10-mer contexts are shown here for ease of viewing. The strong correlation
(Pearson's correlation coefficient = 0.91) between IPDs in identical contexts assayed from completely independent sequencing
runs of different species demonstrate that the context effects are highly consistent between experiments. (C) Graphical representation of the CRF model. The 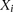 variables represent the hidden modification states for site i, while the
variables represent the hidden modification states for site i, while the 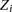 represent the observed IPD values for site i that inform on the modification status of the site. In this model we are considering interactions between the incorporation
site,
represent the observed IPD values for site i that inform on the modification status of the site. In this model we are considering interactions between the incorporation
site, 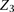 , and the two nearest neighboring sites on each side of
, and the two nearest neighboring sites on each side of 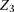 . The edges between the
. The edges between the 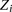 variables indicate there can be interactions between the local sites, with the
variables indicate there can be interactions between the local sites, with the 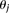 parameters representing the degree of interaction among the nodes. The
parameters representing the degree of interaction among the nodes. The 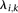 parameters represent the exponential rates for the two possible rate classes at each position i (
parameters represent the exponential rates for the two possible rate classes at each position i (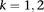 ), while the
), while the 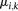 parameters represent the proportion of molecules in state k at position i (with
parameters represent the proportion of molecules in state k at position i (with 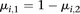 ).
).











