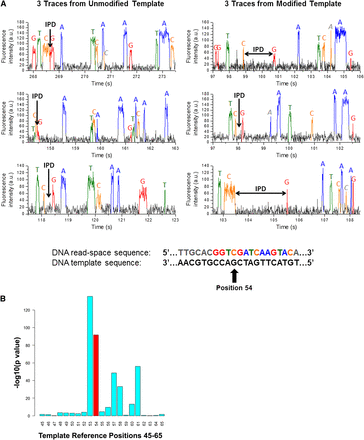
Reproducible variation in interpulse durations as a surrogate for chemical modifications to nucleotide bases. (A) Sample traces of six DNA molecules, three in which the DNA template contains a single 8-oxoG modification (right three traces), and the other identical but with no 8-oxoG modification (left three traces). While the IPD is observed to vary significantly even within the same modification state (a consequence of the exponential nature of the IPD), in the case of the 8-oxoG residue the IPDs are seen to be generally longer than the IPDs of the unmodified G residue. (B) After examining hundreds of molecules in which the G residue was modified versus unmodified, the consistent lengthening of the mean IPD for the modified G residue compared with the mean IPD for the unmodified G residue becomes statistically significant (red bar). The effect of the 8-oxoG modification to the G residue is also seen to affect the IPDs of the neighboring bases in a statistically significant way. In this case, the P-value indicated at each position was computed using the Mann-Whitney test.











