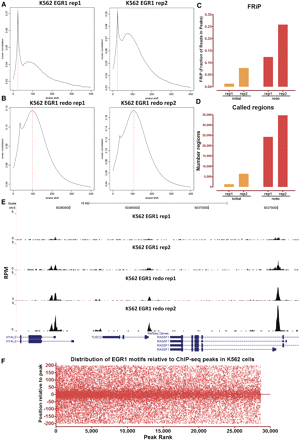
Quality control of ChIP-seq data sets in practice. EGR1 ChIP-seq was performed in K562 cells in two replicates. ChIP enriched regions were identified using MACS. However, the cross-correlation plot profiles (A) indicated that both IPs were suboptimal, with one being unacceptable. In agreement with this judgment, ChIP enrichment (C) and peak number (D) also indicated failure. The ChIP-seq assays were repeated (B), with all quality control metrics improving significantly (B,D), and many additional EGR1 peaks were identified as a result. (E) Representative browser snapshot of the four EGR1 ChIP-seq experiments, showing the much stronger peaks obtained with the second set of replicates. (F) Distribution of EGR1 motifs relative to the bioinformatically defined peak position of EGR1-occupied regions derived from ChIP-seq data in K562 cells. Regions are ranked by their confidence scores as called by SPP.











