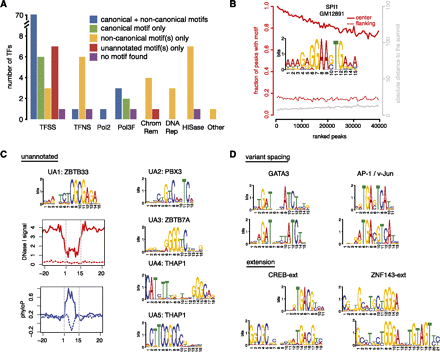
De novo discovery of sequence motifs. (A) Statistics of motif discovery among 119 TFs, classified into 87 Pol II-associated sequence-specific TFs (TFSS), eight general Pol II-associated, non-sequence-specific TFs (TFNS), Pol II (Pol2), six Pol III components and Pol III-associated TFs (Pol3F), five ATP-dependent chromatin complexes (ChromRem), three TFs involved in DNA repair (DNARep), eight histone modification complexes (HISase), and one cyclin kinase associated with transcription (Other). The TATA box binding protein (TBP) is included in the TFNS category and its canonical motif is TATA, corresponding to the blue bar. (B) Example result for SPI1 in GM12891 cells illustrating the percentage of peaks with the motif (left, y-axis in red) and distribution of absolute distances of the closer edge of motif sites relative to the peak summit (right, y-axis in gray), plotted against ranks of peaks (ranked by ChIP-seq signal). (C) Five previously unannotated motifs that are likely to be canonical motifs of four sequence-specific TFs. Also shown are DNase I footprint and sequence conservation profiles around the sites of UA1 (likely the canonical motif of ZBTB33). Motif sites in ChIP-seq peaks (solid lines) were compared with motif sites outside peaks (dashed lines). DNase I and ChIP-seq data were both from K562 cells. Sequence conservation was computed using phyloP (Pollard et al. 2010). (D) Motifs with variant spacing and extensions.











