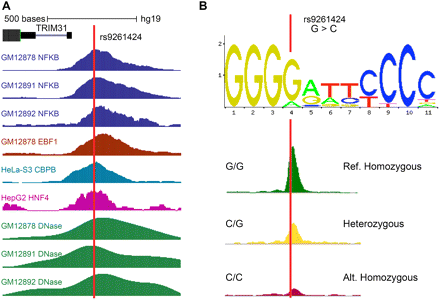
A SNV (rs9261424) overlapping many regulatory features. (A) This SNV falls within peak regions for many ChIP-seq factors as well as DNase-seq peaks from multiple cell lines. (B) The same SNV overlaps a motif match to the NFKB motif and has been shown to alter binding. The signal tracks represent ChIP-seq peaks of NFKB at the SNV site for three individuals: homozygous to reference allele (G), heterozygous, and homozygous to alternate allele (C) (Kasowski et al. 2010).











