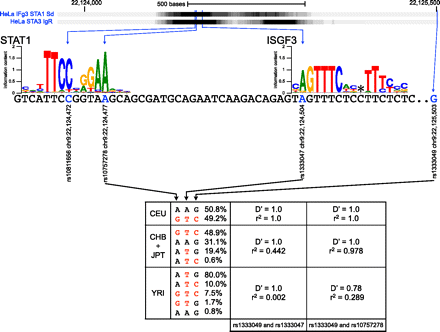
Functional information and linkage disequilibrium patterns support the implication of rs1333047 in coronary artery disease. Functional data (ChIP-seq) generated by the ENCODE Consortium show evidence of STAT1 binding in the 9p21 region associated with coronary artery disease. rs10757278 and rs1333047 are both located in the peak, whereas rs1333049 is a tag SNP that does not overlap any functional region in RegulomeDB. rs10757278 is part of a regulatory motif for STAT1 binding, and rs1333049 is part of a regulatory motif for ISGF3 binding. (*) The location at which a gap is inserted into the motif to handle variable linker length. Haplotype frequency and linkage disequilibrium data from the different HapMap2 populations show that all three SNPs are in perfect linkage disequilibrium in the CEU population, but not the CHB and JPT populations. In the YRI population, the frequency of the A allele at rs1333047 is only 0.8%. Risk alleles for all SNPs are determined using the haplotype associated with coronary artery disease in the CEU population (red). There is an absence of linkage disequilibrium between rs1333047 and rs1333049 in YRI, and the association between rs1333049 and rs10757278 and coronary artery disease has not been replicated in populations of African descent.











