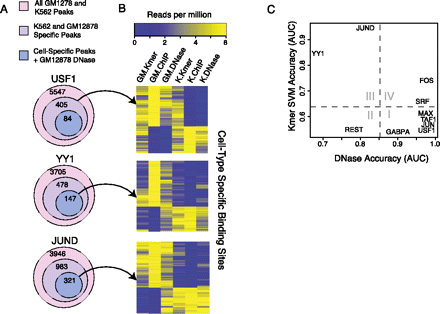
Cell-type–specific sequence models can predict cell-type–specific binding at loci that are DNase accessible in both cell lines. (A) The number of binding sites, cell-type–exclusive binding sites, and exclusive binding sites that are DNase accessible in GM12878. (B) Cell-type–exclusive binding sites can be explained by cell-type–specific sequence preferences when a binding site is accessible in both cell lines. Cell-type–exclusive binding sites for USF1, YY1, and JUND are shown. For USF1, all GM12878- and K562-exclusive binding sites are shown, and DNase accessibility is able to explain cell-type–exclusive binding. In contrast, for JUND and YY1, there are cell-type–exclusive binding sites in GM12878 and K562 that are DNase accessible in both cell lines, and only these examples are plotted in the middle and bottom heatmaps. For these examples, the cell-type–specific SVM sequence scores can explain the cell-type–specific binding. (C) AUC values for the task of discriminating between GM12878-exclusive peaks and K562-exclusive peaks by differential DNase reads (x-axis) or by cell-type–specific SVM sequence scores. For the SVM models, the GM12878- and K562-specific models were each used to discriminate between GM12878- and K562-exclusive binding sites, and the mean AUC over both models was reported. Binding site sequences used in training the models were held out of test sets for this evaluation. For most TFs, the cell-type–exclusive binding sites are well-predicted by differential DNase accessibility (I, IV). For REST, DNase is not predictive in general and the SVM models are consistent between the two cell lines (II). For JUND and YY1 (III), DNase is not predictive of cell-type–exclusive binding, as many sites are DNase accessible in both cell lines; however, the cell-type–specific peaks tend to have different underlying k-mer sequences, enabling accurate discrimination by cell-type–specific SVM sequence models.











