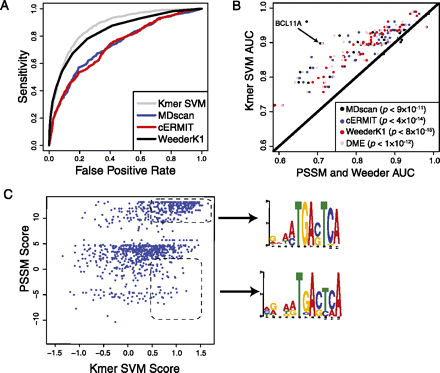
SVM sequence models better predict binding sites than traditional motif approaches. (A) The accuracy of our method is assessed by the area under the ROC curve, which provides a natural trade-off between false positives (x-axis) and sensitivity (y-axis). The ROC curve is shown for discriminating BCL11A ChIP-seq peaks from nonpeaks using four approaches: k-mer SVM, MDscan, cERMIT, and Weeder. (B) The accuracy (AUC) of k-mer SVM models (y-axis) is compared against motif-based algorithms (MDscan, cERMIT, DME, and Weeder; x-axis) for discriminating ChIP-seq peaks from flanking regions. We used training and test sets taken from the same experiment; only accuracy on the test set is shown. Results for transcription factors with multiple ChIP-seq experiments for replicates and cell types were averaged. The SVM models are significantly more accurate than each of the alternative methods (P-values inset and color-coded for each method). (C) The k-mer SVM model is able to learn degenerate motifs. We show the k-mer SVM scores (y-axis) versus the cERMIT motif score (x-axis) for binding sites of BCL11A in GM12878. Example binding sites that are detected by the SVM but receive low scores by the motif are enriched for a more degenerate motif instance, as found by MEME.











