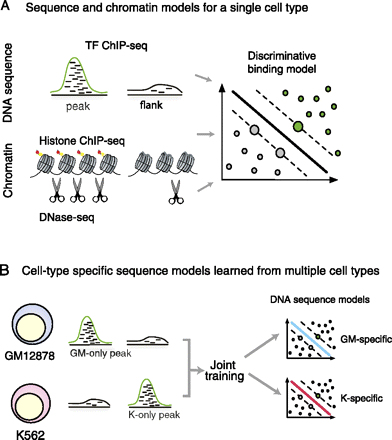
Schematic of models to predict transcription factor occupancy from sequence and chromatin. (A) We developed DNA sequence and chromatin models based on flexible k-mer patterns and spatial organization of histone modifications and DNase accessibility. The models were trained to discriminate between regulatory ChIP-seq peaks and flanking regions within a single cell type using a support vector machine. (B) To study cell-type–specific DNA sequence preferences, we simultaneously train on binding site data from two cell types. This allowed us to jointly learn the cell-type–specific preferences (top and bottom).











