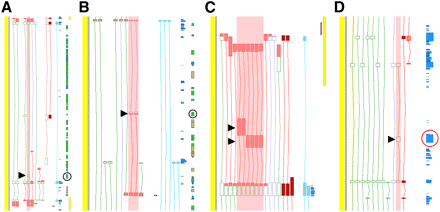
Function of newly identified internal exons of coding loci. Screenshots of gene annotation taken from the Zmap annotation interface. ORF exons of protein-coding models (open green boxes); UTR exons (filled red boxes); variants predicted to be subject to NMD have an ORF represented as open purple boxes. (A–D) Both pre-existing manual annotation and models incorporating model exons identified in this study (highlighted by red shading) are shown. (Black arrowheads) Novel exons; (black circles) overlapping repeat elements; (red circles) overlapping blocks of high cross-species conservation. (A) The BCL2-associated agonist of cell death (BAD) locus. A model incorporating the novel exon is predicted to encode an ORF that breaks the Pfam domain present in other protein-coding models at the locus (Bcl-2_BAD, PF10514) but introduces another domain (GVQW, PF13900) found in caspases, a family of proteins essential for apoptosis, the pathway regulated by the Bcl-2_BAD domain. The novel exon shows no cross-species conservation and overlaps a SINE. (B) The nuclear receptor subfamily 1, group H, member 4 (NR1H4) locus. The two highlighted variants that include the novel exon are both predicted to be subject to NMD and only differ at a small shift in their splice acceptors. Both transcripts are highly liver-specific, and the novel exon was not identified in any other tissue investigated; again there is no cross-species conservation and the novel exon overlaps a SINE. (C) The enoyl-CoA delta isomerase 2 (ECI2) locus. Here two novel exons were identified in the 5′-UTR region of the locus. Although there are only two novel exons, alternative splice donor and acceptor sites in flanking exons suggest many different intron combinations that expand the transcripts repertoire. Neither novel exon overlaps a repeat element or region of cross-species conservation. (D) The KIAA0528 locus. A novel coding transcript incorporating a novel exon remains in-frame relative to other coding transcripts at the locus. The novel exon overlaps a region of exceptionally high conservation as indicated by a peak in the phastCons (44 mammals) track (in blue, circled). The alignment of this exon with other vertebrates' genomes is shown in Supplemental Figure S3.











