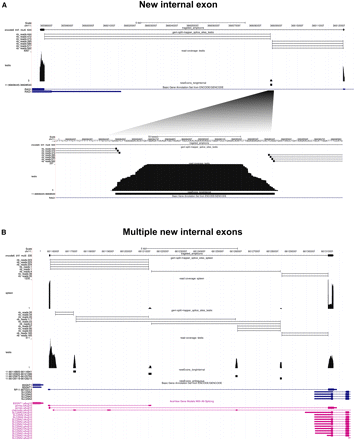
Examples of newly identified internal exons. The two panels show views from the UCSC Genome Browser. The tracks from top to bottom show the scale (black), coordinates (black), the tested GENCODE model (black boxes joined by thin black lines), the introns predicted using the GEM split-mapper, and unmapped reads from the indicated tissue (black ticks joined by thin black lines; number of split-reads for each predicted introns are indicated on the left; see Methods), the RT-PCR-seq sequence reads coverage from the indicated tissue (black; scale on the left), the newly exons identified (black boxes; coordinates are indicated on the left), the ENCODE/GENCODE annotated models (blue or green boxes [exons] joined by thin blue or green lines, respectively), and Aceview predictions using RNA-seq (magenta boxes [exons] joined by thin magenta lines) (Thierry-Mieg and Thierry-Mieg 2006). Identification of a novel internal exon in testis (A) and four novel internal exons and three novel transcript isoforms in spleen and testis (note that some, but not all four, novel exons are supported by RNA-seq) (B).











